Veterinary Laboratory Investigation and Response Network
- Our Mission
- We Respond to Animal Illnesses Potentially Caused by Foods or Drugs
- Vet-LIRN Resources for Animal Owners and Veterinarians
- Fiscal Year 2023 Highlights
- Tracking Antimicrobial Resistance in Bacteria from Sick Animals
- Vet-LIRN Laboratory Funding
- Ensuring Accurate Results
- Veterinarians, Want to Learn More?
- Preparing for and Responding to Emergencies
- Network Laboratory Methods
- Publications
Our Mission
To advance the CVM mission of protecting human and animal health by coordinating a network of veterinary diagnostic laboratories.
Contact Vet-LIRN: vet-lirn@fda.hhs.gov.
We Respond to Animal Illnesses Potentially Caused by Foods or Drugs
Is your animal sick? Do you think it was the food? Or a drug?
- Safety Reporting Portal
- How to Report a Pet Food Complaint
- Reporting Problems with Horse or other Livestock Feed/Food
- Information for Veterinarians on Reporting Suspected Animal Food Issues
Figure 1. What Happens During a Consumer Complaint Response?
We respond to potential animal food issues, including performing non-regulatory testing (Figure 1).
We are an important part of the food safety team at CVM.
Learn more about some of our cases:
- Highly Pathogenic Avian Influenza in Poultry-Based Raw Pet Food
- Response to Influx of Adverse Event Reports Related to Single Pet Food Brand
- Elevated Vitamin D in Commercial Dog Food
- Clostridium botulinum in Alfalfa Cubes
- Campylobacter Outbreak in Puppies
- Aflatoxin Recall
- Pig Ear Outbreak
- Dilated Cardiomyopathy in Some Cases of Pet Heart Disease
- Jerky Pet Treats
Vet-LIRN Resources for Animal Owners and Veterinarians
- Vet-LIRN Network Procedures for Veterinarians
- Vet-LIRN Network Procedures for Owners (En Español)
- Vet-LIRN Network Procedures for Laboratories
- Vet-LIRN SARS-CoV-2 Supplemental Necropsy Sample Inventory Checklist
- Pet Food Safety (CDC)
2024 Highlights
For more information, visit Vet-LIRN Fiscal Year 2024 Accomplishment Highlights.
Tracking Antimicrobial Resistance in Bacteria from Sick Animals
Why track resistance in bacteria?
Antimicrobial resistance is an important public health issue because if bacteria become antibiotic-resistant, many infections become more difficult to treat. In March 2015, The first National Action Plan for Combating Antibiotic-Resistant Bacteria (CARB) was released to guide the government, public heath, healthcare, and veterinary partners in addressing antimicrobial resistance. In 2020, the second CARB plan was released. The 2020 plan builds on the 2015 plan and presents coordinated, strategic actions that the United States Government will take from 2020-2025 by expanding evidence-based activities that have been shown to reduce antibiotic resistance. It also aligns with CVM’s goals to enhance monitoring of animal pathogen antimicrobial resistance as a part of the CVM’s action plan Supporting Antimicrobial Stewardship in Veterinary Settings, Goals for Fiscal Years 2024-2028. As part of this plan, Vet-LIRN was tasked with developing, expanding, and maintaining antimicrobial susceptibility testing (AST) and whole-genome sequencing (WGS) testing of veterinary pathogens isolated at veterinary diagnostic laboratories. To successfully monitor the antimicrobial susceptibility of bacterial pathogens, it is vital that veterinary diagnostic laboratories be incorporated into the nation’s other AMR monitoring activities. Vet-LIRN is committed to being a partner in this effort.
Vet-LIRN Antimicrobial Resistance Monitoring Program Background and Progress
- During 2017-2018, Vet-LIRN coordinated a two-year pilot project to evaluate the feasibility of using Vet-LIRN veterinary diagnostic laboratories to monitor the antimicrobial susceptibility of three veterinary pathogens: Escherichia coli and Staphylococcus pseudintermedius in dogs and Salmonella enterica in any animal host. Twenty Vet-LIRN Source diagnostic laboratories collected isolates and tested their antimicrobial susceptibility using Clinical and Laboratory Standards Institute (CLSI) methods. Approximately 5,000 isolates from clinically sick animals were collected and tested. WGS laboratories sequenced a subset of the isolates submitted by their Source labs and uploaded all sequences to National Center for Biotechnology Information (NCBI) through the GenomeTrakr program. Additional information about each pathogen (the organ it came from, the animal species, which part of the country) was reported. A publication summarizing the 2017 findings is available.
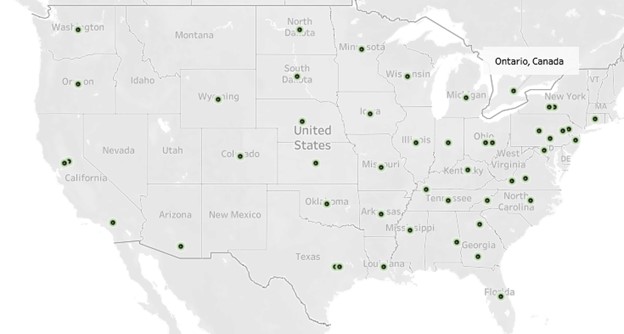
In 2018-2019 additional labs began collecting and sequencing isolates. As of 2024, there are 30 Source laboratories collecting isolates (25 labs in U.S., and 5 labs in Canada) and six WGS laboratories sequencing the isolates. An overview of the general AMR Monitoring plan procedures is provided in Figure 2. More information about participating Source and WGS laboratories is provided in the interactive map of Vet-LIRN AMR Monitoring Program participating laboratories.
Figure 2. Vet-LIRN AMR Monitoring Program: General Plan
Interactive Map of Vet-LIRN AMR Monitoring Program Participating Laboratories (Excel version)
- As of September 2025, Vet-LIRN Source labs collected AST data for more than 30,000 animal pathogen isolates. An overview of the animal pathogens monitored is provided in Table 1. Antimicrobial susceptibility testing data associated with these isolates are publicly available in an AMR Database (Excel version) which contains the phenotypic testing data for all isolates collected from 2017 to 2024.
- Guidance to AMR Database viewers: antimicrobial resistance is extremely complex and driven by many factors. In general, it is difficult to draw meaningful conclusions by comparing just one year to another. Instead, it is best to look for patterns that emerge over several years.
- Persons who use these data should cite the Veterinary Laboratory Investigation and Response Network (Vet-LIRN) as the source of the original data. The data in this database is not confidential. Suggested citation: Food and Drug Administration (FDA). Vet-LIRN. Laurel, MD: U.S. Department of Health and Human Services. Available from URL: https://www.fda.gov/animal-veterinary/science-research/veterinary-laboratory-investigation-and-response-network. Accessed MM/DD/YYYY.
- People using assistive technology such as a screen reader may not be able to fully access all permutations of the information available in the interactive resistance displays. To gain alternate access to the information, you may download spreadsheets containing the source data. People with disabilities who require further assistance may contact Vet-LIRN@fda.hhs.gov.
- The data provides a snapshot of the susceptibility of pathogens being cultured at referral veterinary laboratories.
- Sequencing data are released in real-time as whole-genome sequencing is conducted. More than 8,500 isolates were sequenced. Antimicrobial susceptibility testing data associated with these isolates are also publicly available.
- Vet-LIRN partners with the National Antimicrobial Resistance Monitoring System (NARMS) to make the data public (Animal Pathogen AMR Data). This animal pathogen data is reported in conjunction with the National Animal Health Laboratory Network (NAHLN).
Table 1: List of animal pathogens and animal hosts
| Bacterial pathogen | Animal host | AST/WGS testing (yes/no) |
|---|---|---|
| Salmonella enterica | Any animal host | Yes/Yes |
| Escherichia coli | Dogs | Yes/Yes |
| Staphylococcus pseudintermedius | Dogs | Yes/Yes |
| Staphylococcus aureus | Pigs, cattle, sheep, goats, poultry, dogs, cats, and horses | Yes/Yes |
| S. schleiferi and/or S. coagulans | Dogs & cats | Yes /Yes |
| (a) Klebsiella pneumoniae/variicola (b) Pseudomonas aeruginosa (c) Enterococcus faecalis or faecium | (a) Any animal host (b) Dogs, horses (c) Dogs, cats, poultry | Yes/Yes |
| (d) Enterobacter cloacae complex (e) Acinetobacter baumannii complex | (d) and (e) Dogs & cats | Yes/Yes |
Report AST data for confirmed Carbapenem-resistant Enterobacterales and/or Carbapenem-resistant Organisms
| Any animal host | Yes/Yes |
| (f) Campylobacter coli (f) Campylobacter jejuni (f) Campylobacter fetus | (f) Swine, poultry, cattle, small ruminants, dogs, cats | No/Yes |
| Fish pathogens: any bacterial pathogen | Fish | No/Yes |
Promoting Antimicrobial Stewardship
Along with tracking antimicrobial resistance, Vet-LIRN is working to promote antimicrobial stewardship in veterinary medicine, in line with CVM's Initiatives promoting antimicrobial stewardship. As described above, antimicrobial resistance is an important public health issue, and use of antimicrobial drugs can contribute to the development of antimicrobial resistant bacteria. Antimicrobial stewardship involves using antimicrobials appropriately and only when necessary.
Vet-LIRN supports antimicrobial stewardship efforts by providing funding to veterinary colleges across the United States to work on several projects including:
- creating collaborative websites and publications with antimicrobial resistance resources,
- generating veterinary hospital stewardship plans,
- developing educational materials for veterinary health professionals (including veterinarians, veterinary diagnostic laboratories, and veterinary students), and
- developing educational materials for animal owners and producers.
These materials consist of website content, fliers, videos, online course modules, and other formats to encourage education and appropriate use of antimicrobials in the veterinary community.
Examples of Stewardship Educational Materials
Carbapenem-Resistant Enterobacterales (CRE) - Website
Antibiotic Resistant Bacteria in Companion Animals - Fliers
One Health Approach for Reporting Veterinary Carbapenem-Resistant Enterobacterales and Other Bacteria of Public Health Concern - Volume 29, Number 6—June 2023 - Emerging Infectious Diseases journal - CDC
A Multicenter Evaluation of a Metacognitive Framework for Antimicrobial Selection Education - Journal of Veterinary Medical Education
Vet-LIRN Laboratory Funding
Vet-LIRN Cooperative Agreements facilitate participation in Vet-LIRN activities such as consumer complaint response, emergency exercises, proficiency tests, and laboratory accreditation. The agreements also increase the agency’s capability to analyze an increased number of samples in the event of animal food- or drug-related illnesses or other large-scale emergency events that require increased testing of implicated diagnostic or animal food samples. Cooperative agreements allow network laboratories to request additional funds if they are participating in a specific Vet-LIRN project, such as the Antimicrobial Resistance (AMR) Project or if they are conducting whole-genome sequencing (WGS) work, or if their caseload is particularly heavy. Additional funds may also be provided to respond to emerging diseases.
Ensuring Accurate Results
Vet-LIRN collaborates with the FDA’s Human Foods Program (HFP) Division of Food Processing Science and Technology (Moffett Center) and the Institute for Food Safety and Health, Illinois Institute of Technology to conduct Proficiency Tests (PTs) and Interlaboratory Comparison Exercises (ICEs) to ensure FDA receives accurate test results from our network laboratories. Samples are sent to the laboratories and test results are submitted to Vet-LIRN. After data is evaluated, final reports are provided to the laboratories.
Recent Proficiency Tests and Inter-Laboratory Comparison Exercises
- Detecting of avian influenza in milk
Timely to support veterinary diagnostic laboratories’ ability to evaluate the accuracy of their current testing methods. - Detecting Salmonella in dog food
- Detecting an Mycotoxin in dog food
- Whole Genome Sequencing Proficiency Test
- Why is this important? It evaluates network laboratories’ ability to sequence and identify a variety of bacteria isolates (from start to finish) that could cause potential issues with animal foods.
Veterinarians, Want to Learn More?
Vet-LIRN educates veterinarians about how to identify and report suspected animal food issues via webinars and case studies. Vet-LIRN speaks at various conferences and to veterinary interest groups. Additionally, Vet-LIRN can present virtual lectures to veterinary schools to increase awareness among future veterinarians of CVM’s mission and consumer complaint reporting.
Please email Vet-LIRN@fda.hhs.gov if you are interested in a presentation to your institution.
Preparing for and Responding to Emergencies
Vet-LIRN participates in simulated incidents (exercises) and evaluation of emergency preparedness and response activities. Such activities strengthen Vet-LIRN’s ability to establish and initiate strategies to coordinate the roles and responsibilities of veterinary diagnostics laboratories in real-world emergency events. Understanding network laboratory capabilities and establishing routine interactions and exercises with the laboratories is key to any emergency preparedness and response. Vet-LIRN routinely communicates with the following laboratory networks and programs to harmonize and leverage activities and participate in an integrated response to national emergencies:
- Integrated Consortium of Laboratory Networks (ICLN)
- National Animal Health Laboratory Network (NAHLN)
- The Food Emergency Response Network (FERN)
Network Laboratory Methods
Vet-LIRN is working to ensure that detailed protocols and procedures of methods developed from grant funding are publicly available. All protocols and procedures published are available at Vet LIRN.
Publications (Listed past 5 years)
Kiener, S., Smith, E., Singh, N., Nemser, S. M., Hettwer, K., Miller, M. R., Tkachenko, A., Uhlig, S., & Reddy, R. (2025). Determination of limit of detection and relative limit of detection of Salmonella in raw pet food matrices using Salmonella bacteriological analytical manual methods. J Microbiol Methods, 232-234, 107116. https://doi.org/10.1016/j.mimet.2025.107116
Martin, G., Tyson, G. H., Guag, J., Strain, E., & Ceric, O. (2025). Genomic snapshot of Klebsiella spp. isolates from clinically ill animals reveal diverse lineages with limited relatedness to human isolates. BMC Vet Res, 21(1), 458. https://doi.org/10.1186/s12917-025-04686-z
Nemser, S., Ceric, O., Guag, J., Pauley, S., Jones, A., Proia, K., Miller, M., Tkachenko, A., Rostein, D., Hodges, A., Reimschessel, R., Tyson, G. (2025). The Veterinary Laboratory Investigation and Response Network: 15 Years of Promoting Human and Animal Health by Collaborating with the Veterinary Diagnostic Laboratory Community. Journal of Food Protection. (In press)
Pepper, A., Kayastha, S., Miller, M., Guag, J., Tkachenko, A., Allender, M., Terio, K., & Wang, L. (2025). Evaluation of RT-LAMP for SARS-CoV-2 Detection in Animal Feces. Viruses, 17(6), 783. https://www.mdpi.com/1999-4915/17/6/783
Singh, N., Miller, M. R., Nemser, S. M., Tkachenko, A., Uhlig, S., Frost, K., Hettwer, K., Ulaszek, J., Kmet, M., Wang, L., Allender, M. C., & Reddy, R. (2025). Proficiency test of SARS-CoV-2 Omicron variant detection in diagnostics samples by veterinary diagnostic laboratories. Accreditation and Quality Assurance, 30(1), 45-53. https://doi.org/10.1007/s00769-024-01622-w
Tkachenko, A., Chen, Y., Petrey, M., Fritz, S., Walsh, T., Rotstein, D., Miller, M. R., Williams, B., Dark, M., Kmet, M., Reddy, R., Tyson, G., & Nemser, S. M. (2025). A novel proficiency test to assess the animal diagnostic investigation process in identifying an unknown toxicant. Toxicology Reports, 14, 101925. https://doi.org/https://doi.org/10.1016/j.toxrep.2025.101925
Zehr, J. D., Sun, Q., Ceres, K., Merrill, A., Tyson, G. H., Ceric, O., Guag, J., Pauley, S., McQueary, H. C., Sams, K., Reboul, G., Mitchell, P. K., Anderson, R., Franklin-Guild, R., Guarino, C., Cronk, B. D., Burbick, C. R., Wolking, R., Peak, L.,…Goodman, L. B. (2025). Population and pan-genomic analyses of Staphylococcus pseudintermedius identify geographic distinctions in accessory gene content and novel loci associated with AMR. Appl Environ Microbiol, e0001025. https://doi.org/10.1128/aem.00010-25
Ceres K, Zehr JD, Murrell C, Millet JK, Sun Q, McQueary HC, Horton A, Cazer C, Sams K, Reboul G, Andreopoulos WB, Mitchell PK, Anderson R, Franklin-Guild R, Cronk BD, Stanhope BJ, Burbick CR, Wolking R, Peak L, Zhang Y, McDowall R, Krishnamurthy A, Slavic D, Sekhon PK, Tyson GH, Ceric O, Stanhope MJ, Goodman LB. 2024. Evolutionary genomic analyses of canine E. coli infections identify a relic capsular locus associated with resistance to multiple classes of antimicrobials. Appl Environ Microbiol 90:e00354-24.https://doi.org/10.1128/aem.00354-24
Nichols, M., Stapleton, G. S., Rotstein, D. S., Gollarza, L., Adams, J., Caidi, H., . . . Francois Watkins, L. K. (2024). Outbreak of multidrug-resistant Salmonella infections in people linked to pig ear pet treats, United States, 2015-2019: results of a multistate investigation. Lancet Reg Health Am, 34, 100769. doi:10.1016/j.lana.2024.100769
Langston, J., Stump, S., Filigenzi, M., Tkachenko, A., Guag, J., Poppenga, R., & Rumbeiha, W. K. (2024). Extensive evaluation of a new LC-MS/MS method to quantify monofluoroacetate toxin in the kidney. J Anal Toxicol. doi:10.1093/jat/bkae032
Leonardi-Cattolica, A., Kayastha, S., Miller, M., Guag, J., Tkachenko, A., Lowe, J., Allender, M., Terio, K., & Wang, L. (2024). Evaluation of Fecal Sample Pooling for Real-Time RT-PCR Testing SARS-CoV-2 in Animals. Viruses, 16(11). https://doi.org/10.3390/v16111651
Miller MR, Tkachenko A, Guag J, Alexander S, Webb BT, Stenger BLS. Comparative evaluation of assay performance for SARS-CoV-2 detection in animal oral samples, lung homogenates, and phosphate-buffered saline using the TaqPath COVID-19 Combo kit. J Vet Diagn Invest. 2024 Mar;36(2):229-237. doi: 10.1177/10406387241230315. Epub 2024 Feb 16. PMID: 38362609; PMCID: PMC10929630.
Eckstrand, C. D., Torrevillas, B. K., Wolking, R. M., Francis, M., Goodman, L. B., Ceric, O., Alexander, T. L., Snekvik, K. R., & Burbick, C. R. (2024). Genomic characterization of antimicrobial resistance in 61 aquatic bacterial isolates. J Vet Diagn Invest, 10406387241241042. https://doi.org/10.1177/10406387241241042
Kattoor, J. J., Guag, J., Nemser, S. M., & Wilkes, R. P. (2024). Development of ion torrent-based targeted next-generation sequencing panel for identification of animal species in pet foods. Res Vet Sci, 167, 105117. https://doi.org/10.1016/j.rvsc.2023.105117
Miller MR, Braun E, Ip HS, Tyson GH. Domestic and wild animal samples and diagnostic testing for SARS-CoV-2. Vet Q. 2023 Dec;43(1):1-11. doi: 10.1080/01652176.2023.2263864. Epub 2023 Oct 26. PMID: 37779468; PMCID: PMC10614713.
Chen, Y., Lopez, S., Reddy, R. M., Wan, J., Tkachenko, A., Nemser, S. M., Smith, L., & Reimschessel, R. (2023). Validation and interlaboratory comparison of anticoagulant rodenticide analysis in animal livers using ultra-performance liquid chromatography-mass spectrometry. J Vet Diagn Invest, 35(5), 470-483. https://doi.org/10.1177/10406387231178558
Deng, K., Nemser, S. M., Frost, K., Goodman, L. B., Ip, H. S., Killian, M. L., . . . Tyson, G. H. (2023). Successful Detection of Delta and Omicron Variants of SARS-CoV-2 by Veterinary Diagnostic Laboratory Participants in an Interlaboratory Comparison Exercise. The Journal of Applied Laboratory Medicine. doi:10.1093/jalm/jfad018
Francis, K. A., Tkachenko, A., Johnson, J. T., Smith, L. L., Noonan, R. T., Filigenzi, M. S., . . . Romano, M. C. (2023). Comprehensive Evaluation of HPLC-MS/MS Method for Quantitation of Seven Anticoagulant Rodenticides and Dicoumarol in Animal Serum. J Anal Toxicol. doi:10.1093/jat/bkad017
Du, X., Schrunk, D. E., Imerman, P. M., Tahara, J., Tkachenko, A., Guag, J., . . . Rumbeiha, W. K. (2023). Extensive Evaluation of a Method for Quantitative Measurement of Aflatoxins B1 and M1 in Animal Urine Using High-Performance Liquid Chromatography with Fluorescence Detection. J AOAC Int, 106(3), 645-651. doi:10.1093/jaoacint/qsad0342022
Ballash, G. A., Dennis, P. M., Mollenkopf, D. F., Albers, A. L., Robison, T. L., Adams, R. J., . . . Wittum, T. E. (2022). Colonization of White-Tailed Deer (Odocoileus virginianus) from Urban and Suburban Environments with Cephalosporinase- and Carbapenemase-Producing Enterobacterales. Appl Environ Microbiol, 88(13), e0046522. doi:10.1128/aem.00465-22
Deng, K., Uhlig, S., Goodman, L. B., Ip, H. S., Killian, M. L., Nemser, S. M., Tkachenko, A . . . Tyson, G. H. (2022). Second round of an interlaboratory comparison of SARS-CoV2 molecular detection assays used by 45 veterinary diagnostic laboratories in the United States. J Vet Diagn Invest, 34(5), 825-834. doi:10.1177/10406387221115702
Esmaeilishirazifard, E., Usher, L., Trim, C., Denise, H., Sangal, V., Tyson, G. H., . . . Moschos, S. A. (2022). Bacterial Adaptation to Venom in Snakes and Arachnida. Microbiol Spectr, 10(3), e0240821. doi:10.1128/spectrum.02408-21
Harrison, L., Tyson, G. H., Strain, E., Lindsey, R. L., Strockbine, N., Ceric, O., . . . Dessai, U. (2022). Use of Large-Scale Genomics to Identify the Role of Animals and Foods as Potential Sources of Extraintestinal Pathogenic Escherichia coli That Cause Human Illness. Foods, 11(13). doi:10.3390/foods11131975
Mitchell, P. K., Wang, L., Stanhope, B. J., Cronk, B. D., Anderson, R., Mohan, S., . . . Goodman, L. B. (2022). Multi-laboratory evaluation of the Illumina iSeq platform for whole genome sequencing of Salmonella, Escherichia coli and Listeria. Microb Genom, 8(2). doi:10.1099/mgen.0.000717
Rotstein, D. S., Peloquin, S., Proia, K., Hart, E., Lee, J., Vyhnal, K. K., . . . Ghai, R. (2022). Investigation of SARS-CoV-2 infection and associated lesions in exotic and companion animals. Vet Pathol, 3009858211067467. doi:10.1177/03009858211067467
Tate, H., Ayers, S., Nyirabahizi, E., Li, C., Borenstein, S., Young, S., . . . McDermott, P. F. (2022). Prevalence of Antimicrobial Resistance in Select Bacteria From Retail Seafood-United States, 2019. Front Microbiol, 13, 928509. doi:10.3389/fmicb.2022.928509
Deng, K., Uhlig, S., Ip, H. S., Lea Killian, M., Goodman, L. B., Nemser, S., . . . Reimschuessel, R. (2021). Interlaboratory comparison of SARS-CoV2 molecular detection assays in use by U.S. veterinary diagnostic laboratories. J Vet Diagn Invest, 33(6), 1039-1051. doi:10.1177/10406387211029913
Girard, L., Herath, K., Escobar, H., Reimschuessel, R., Ceric, O., & Jayasuriya, H. (2021). Development of UHPLC/Q-TOF Analysis Method to Screen Glycerin for Direct Detection of Process Contaminants 3-Monochloropropane-1,2-diol Esters (3-MCPDEs) and Glycidyl Esters (GEs). Molecules, 26(9). doi:10.3390/molecules26092449
Nemser, S., Lindemann, S., Chen, Y., Lopez, S., Pickens, S., Ulaszek, J., . . . Reddy, R. (2021). A review of proficiency exercises offered by the Veterinary Laboratory Investigation and Response Network (Vet-LIRN) and Moffett Proficiency Testing Laboratory from 2012 to 2018. Accreditation and Quality Assurance, 26(3), 143-156. doi:10.1007/s00769-021-01471-x
Rotstein, D., Jones, J. L., Buchweitz, J., Refsal, K. R., Wilson, R., Yanes, E. G., . . . Reimschuessel, R. (2021). Pet Food-Associated Dietary Exogenous Thyrotoxicosis: Retrospective Study (2016-2018) and Clinical Considerations. Top Companion Anim Med, 43, 100521. doi:10.1016/j.tcam.2021.100521
Peloquin, S. K., Rotstein, D. S., Jones, J. L., Guag, J., Carey, L., Palmer, L. A., . . . Reimschuessel, R. (2021). Presumed Choline Chloride Toxicosis in Cats With Positive Ethylene Glycol Tests After Consuming a Recalled Cat Food. Top Companion Anim Med, 44, 100548. doi:10.1016/j.tcam.2021.100548
Taghvaei, M., Tonyali, B., Sommers, C., Ceric, O., Linghu, Z., Smith, J. S., & Yucel, U. (2021). Formation kinetics of radiolytic lipid products in model food–lipid systems with gamma irradiation. Journal of the American Oil Chemists' Society, 98(7), 737-746. doi:https://doi.org/10.1002/aocs.12513
Tkachenko, A., Benson, K., Mostrom, M., Guag, J., Reimschuessel, R., & Webb, B. (2021). Extensive evaluation via blinded testing of an UHPLC-MS/MS method for quantitation of ten ergot alkaloids in rye and wheat grains. J AOAC Int. doi:10.1093/jaoacint/qsaa173
Tyson, G. H., Ceric, O., Guag, J., Nemser, S., Borenstein, S., Slavic, D., . . . Reimschuessel, R. (2021). Genomics accurately predicts antimicrobial resistance in Staphylococcus pseudintermedius collected as part of Vet-LIRN resistance monitoring. Veterinary Microbiology, 254, 109006. doi:https://doi.org/10.1016/j.vetmic.2021.109006
Vudathala, D., Cummings, M., Tkachenko, A., Guag, J., Reimschuessel, R., & Murphy, L. (2021). A Lateral Flow Method for Aflatoxin B1 in Dry Dog Food: An Inter-Laboratory Trial. J AOAC Int. doi:10.1093/jaoacint/qsaa175
Publications (Vet-LIRN funded) (Listed past 3 years)
Ardalan, M., Cool, K., Gaudreault, N. N., Bold, D., Mannix, A., Hanzlicek, G. A., Richt, J. A., & Pogranichniy, R. M. (2024). Cattle, sheep, and goat humoral immune responses against SARS-CoV-2. Vet Anim Sci, 26, 100408. https://doi.org/10.1016/j.vas.2024.100408
Ardalan, M., Cool, K., Gaudreault, N. N., Bold, D., Rojas, C., Mannix, A., Seetahal, J., Richt, J. A., & Pogranichniy, R. M. (2024). Bison, Elk, and Other Captive Wildlife Species Humoral Immune Responses against SARS-CoV-2. Animals (Basel), 14(19). https://doi.org/10.3390/ani14192829
Lopez, K. P., Cool, K. R., Bold, D., Gaudreault, N. N., Roberts, B. A., Maag, E., Richt, J. A., & Pogranichniy, R. M. (2025). Detection of SARS-CoV-2- specific antibodies in domestic cats using different ELISA tests. J Virol Methods, 333, 115099. https://doi.org/10.1016/j.jviromet.2024.115099
Bacon, R. L., Hodo, C. L., Wu, J., Welch, S., Nickodem, C., Vinasco, J., Threadgill, D., Gray, S. B., Norman, K. N., & Lawhon, S. D. (2024). Diversity of Campylobacter spp. circulating in a rhesus macaque (Macaca mulatta) breeding colony using culture and molecular methods. mSphere, 9(11), e0056024. https://doi.org/10.1128/msphere.00560-24
Hooser, S. B. (2024). Investigative and Diagnostic Toxicology and Feed-Related Outbreaks. Veterinary Clinics of North America: Equine Practice, 40(1), 1-10. https://doi.org/10.1016/j.cveq.2023.12.001
Langston, J., Stump, S., Filigenzi, M., Tkachenko, A., Guag, J., Poppenga, R., & Rumbeiha, W. K. (2024). Extensive evaluation of a new LC-MS/MS method to quantify monofluoroacetate toxin in the kidney. J Anal Toxicol. doi:10.1093/jat/bkae032
Comparative evaluation of assay performance for SARS-CoV-2 detection in animal oral samples, lung homogenates, and phosphate-buffered saline using the TaqPath COVID-19 Combo kit - Megan R. Miller, Andriy Tkachenko, Jake Guag, Stacey Alexander, Brett T. Webb, Brianna L. S. Stenger, 2024 (sagepub.com)
Thieulent CJ, Carossino M, Peak L, Wolfson W, Balasuriya UBR. Multiplex One-Step RT-qPCR Assays for Simultaneous Detection of SARS-CoV-2 and Other Enteric Viruses of Dogs and Cats. Viruses. 2023 Sep 7;15(9):1890. doi: 10.3390/v15091890. PMID: 37766296; PMCID: PMC10534472.
Thieulent CJ, Carossino M, Peak L, Strother K, Wolfson W, Balasuriya UBR. Development and Validation of a Panel of One-Step Four-Plex qPCR/RT-qPCR Assays for Simultaneous Detection of SARS-CoV-2 and Other Pathogens Associated with Canine Infectious Respiratory Disease Complex. Viruses. 2023 Sep 5;15(9):1881. doi: 10.3390/v15091881. PMID: 37766287; PMCID: PMC10535912.
Meisner, J., Baszler, T. V., Kuehl, K. E., Ramirez, V., Baines, A., Frisbie, L. A., . . . Rabinowitz, P. M. (2022). Household Transmission of SARS-CoV-2 from Humans to Pets, Washington and Idaho, USA. Emerg Infect Dis, 28(12), 2425-2434. doi:10.3201/eid2812.220215