2021 FDA Science Forum
Species Level Identification from Complex Mixtures Using Colony Mass Spectrometry
- Authors:
- Center:
-
Contributing OfficeCenter for Biologics Evaluation and Research
Abstract
Background
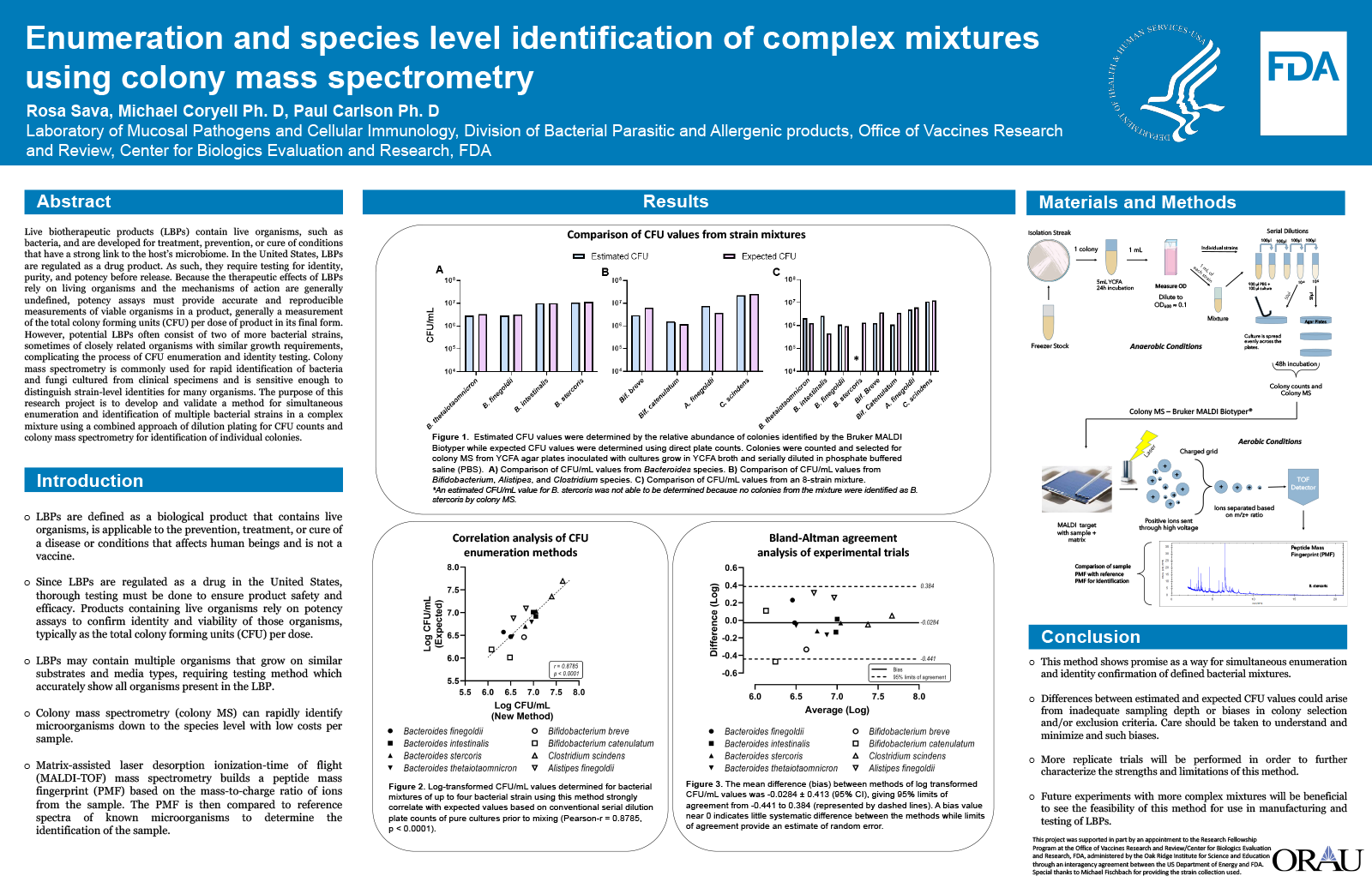
Live biotherapeutic products (LBPs) contain live organisms, such as bacteria, and are developed for treatment, prevention, or cure of conditions that have a strong link to the host’s microbiome. In the United States, LBPs are regulated as a drug product. As such, they require testing for identity, purity, and potency before release. Because the therapeutic effects of LBPs rely on living organisms and the mechanisms of action are generally undefined, potency assays must provide accurate and reproducible measurements of viable organisms in a product, generally a measurement of the total colony forming units (CFU) per dose of product in its final form. However, potential LBPs often consist of two of more bacterial strains, sometimes of closely related organisms with similar growth requirements, complicating the process of CFU enumeration and identity testing. Colony mass spectrometry is commonly used for rapid identification of bacteria and fungi cultured from clinical specimens and is sensitive enough to distinguish strain-level identities for many organisms. The purpose of this research project is to develop and validate a method for simultaneous enumeration and identification of multiple bacterial strains in a complex mixture using a combined approach of dilution plating for CFU counts and colony mass spectrometry for identification of individual colonies.
Methodology
Bacterial strains were isolated and grown in YCFA media before creating a mixture containing equal parts of each strain. A dilution series was then performed on the individual and mixed cultures to enumerate CFUs and isolated colonies were selected for identification using colony mass spectrometry. CFU/mL of each strain was estimated by multiplying the relative abundance of the colony by the total CFU/mL in the mixture. There was a high degree of agreement between the expected and empirical CFU measurements in bacterial mixtures.
Conclusion
Coupled with traditional CFU enumeration, colony mass spectrometry can be used to rapidly identify the intended bacterial strains in a complex mixture of organisms. This method represents a potential tool for both potency and identity testing of defined bacterial products with potential applications in testing of LBPs as well as other bacterial consortia used in industry, agriculture, and veterinary medicine.