2023 FDA Science Forum
RIPS: Routine Intuitive Pathogen surveillance with whole genome sequence data
- Authors:
- Center:
-
Contributing OfficeCenter for Food Safety and Applied Nutrition
Abstract
Background:
Genomes from bacterial pathogens, provided by whole genome sequence (WGS) collection programs, are monitored for emerging outbreaks by finding matches between new and previously submitted data originating from foods, environments, and patients. The FDA CFSAN’s Biostatistics and Bioinformatics Staff (BBS) uses specialized tools for WGS surveillance, which limits their use. Here, we present an accessible tool for routine WGS surveillance (RIPS – Routine Intuitive Pathogen Surveillance).
Purpose:
RIPS makes routine surveillance of whole genome sequence data accessible to a wide range of end-users, democratizing the detection of emerging outbreaks while they are small, which may help to reduce the numbers of illnesses. RIPS provides a means to apply uniform and objective criteria that define emerging outbreaks. The criteria are tunable, which allows the user to measure the outcome of different settings to ensure optimal performance.
Methodology:
RIPS was written with the Shiny package in R. It downloads Rapid Reports data from the Pathogen Detection database maintained by the National Center for Biotechnology Information. Results of a preliminary analysis that compares newly submitted sequence data to all other sequences in the database are further filtered using metrics, such as the numbers of sequences for bacteria collected from patients (clinical isolates) within a stated timeframe and the genomic distances between clinical and non-clinical isolates. The information is displayed in an easy-to-understand format, along with additional information, such as serotypes and geography.
Results:
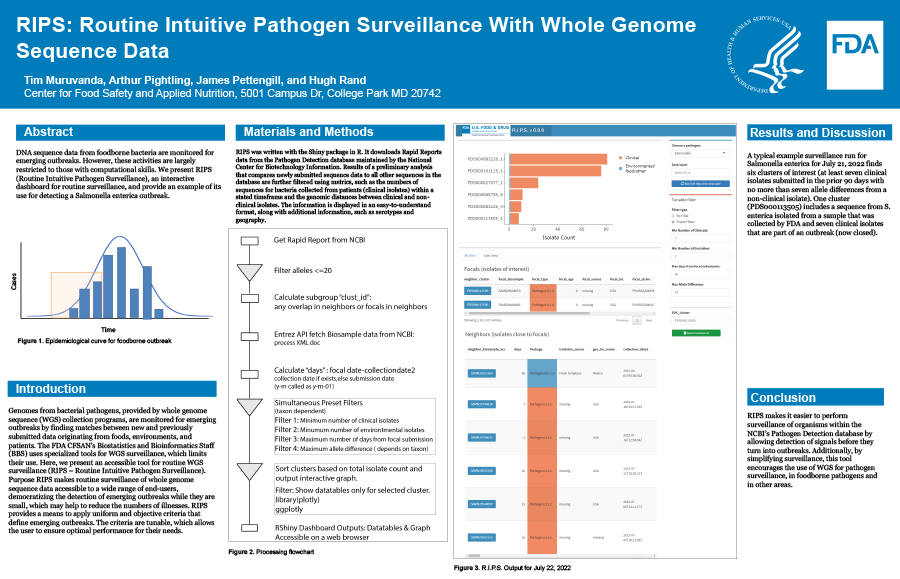
Comparisons with the existing methodology indicate RIPS is fast and accurate. A typical example surveillance run for Salmonella enterica for July 21, 2022 finds six clusters of interest (at least seven clinical isolates submitted in the prior 90 days with no more than seven allele differences from a non-clinical isolate). One cluster (PDS000113505) includes a sequence from S. enterica isolated from a sample that was collected by FDA and seven clinical isolates that are part of an outbreak (now closed).
Conclusion:
RIPS makes it easier to perform surveillance of organisms within the NCBI’s Pathogen Detection database. By simplifying surveillance, this tool encourages the use of WGS for pathogen surveillance, in foodborne pathogens and in other areas.