2023 FDA Science Forum
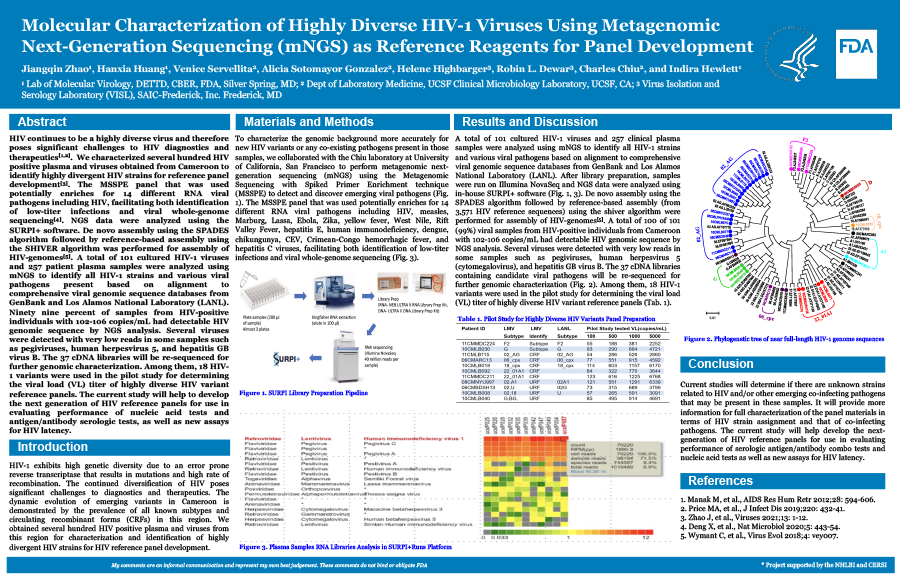
Molecular Characterization of Highly Diverse HIV-1 Viruses Using Metagenomic Next-Generation Sequencing (mNGS) as Reference Reagents for Panel Development
- Authors:
- Center:
-
Contributing OfficeCenter for Biologics Evaluation and Research
Abstract
Background and Purpose
HIV continues to be a highly diverse virus and therefore poses significant challenges to HIV diagnostics and therapeutics. There is a need for ongoing surveillance of emerging HIV variants, assessing impact on diagnostics, and periodic updating of reference panels with globally prevalent diverse strains for non-detection and under-quantitation by some commercial NAT assays. We characterized several hundred HIV positive plasma and viruses obtained from Cameroon to identify highly divergent HIV strains for reference panel development. Methodology In collaboration with Dr. Chiu’s Lab we performed mNGS using the Metagenomic Sequencing with Spiked Primer Enrichment technique (MSSPE) to detect and discover emerging viral pathogens. The MSSPE panel that was used potentially enriches for 14 different RNA viral pathogens including HIV, facilitating both identification of low-titer infections and viral whole-genome sequencing. NGS data were analyzed using the SURPI+ software. De novo assembly using the SPADES algorithm followed by reference-based assembly using the SHIVER algorithm was performed for assembly of HIV-1 genomes. Results A total of 101 cultured HIV-1 viruses and 257 patient plasma samples were analyzed using mNGS to identify all HIV-1 strains and various viral pathogens present based on alignment to comprehensive viral genomic sequence databases from GenBank and Los Alamos National Laboratory (LANL). Ninety nine percent of samples from HIV-positive individuals with 102-106 copies/mL had detectable HIV genomic sequence by NGS analysis. Several viruses were detected with very low reads in some samples such as pegiviruses, human herpesvirus 5, and hepatitis GB virus B. The 37 cDNA libraries will be re-sequenced for further genomic characterization. Among them, 18 HIV-1 variants were used in the pilot study for determining the viral load (VL) titer of highly diverse HIV variant reference panels. Conclusions This study identified strains related to HIV and/or other emerging co-infecting pathogens that may be present in these samples and thereby provide in-depth characterization of panel materials in terms of HIV strain assignment and co-infecting pathogens. The current study will help to develop the next generation of HIV reference panels for use in evaluating performance of nucleic acid tests and antigen/antibody serologic tests, as well as new assays for HIV latency. (My comments are an informal communication and represent my own best judgement. These comments do not bind or obligate FDA.)