2021 FDA Science Forum
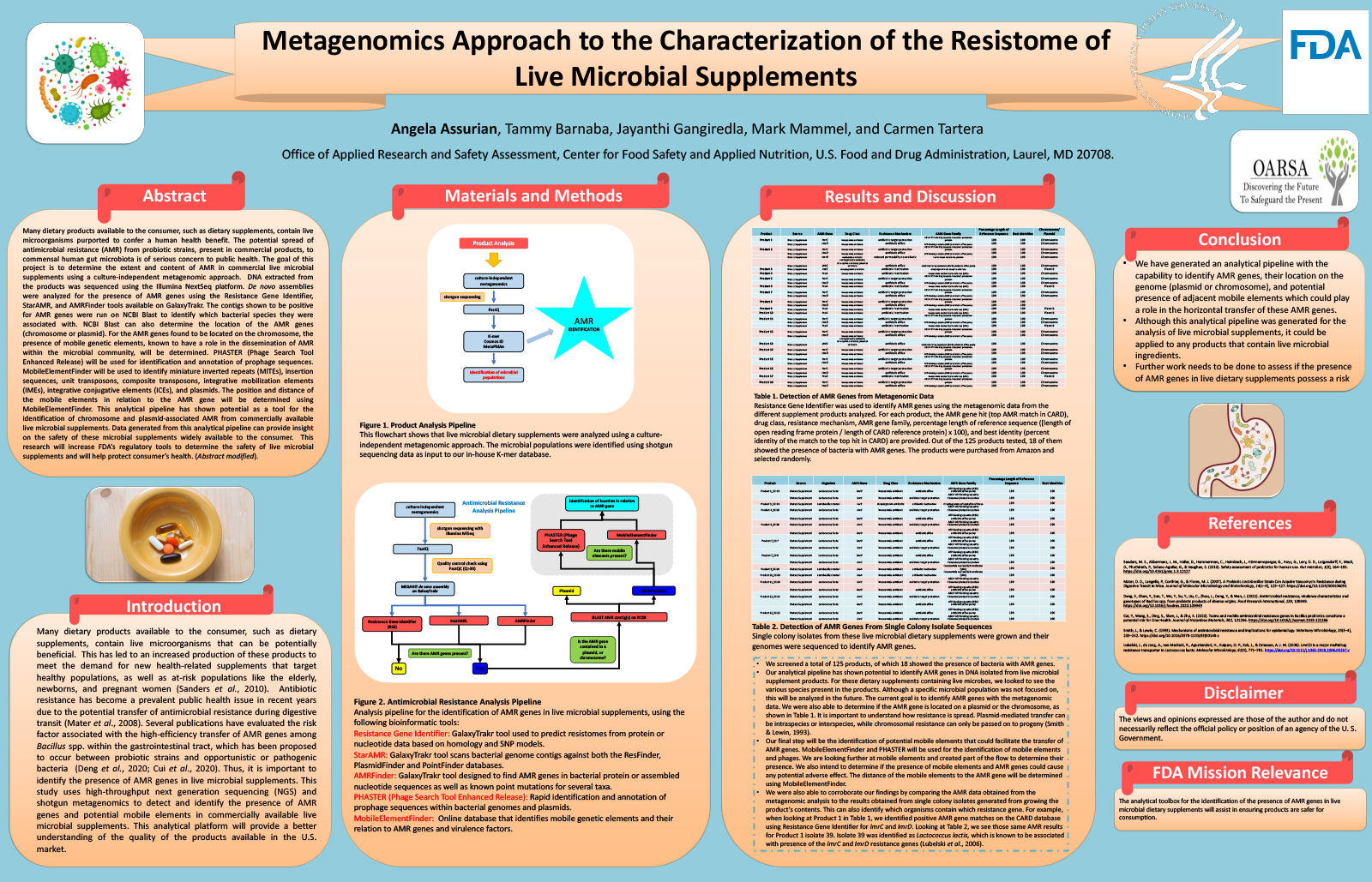
Metagenomics Approach to the Characterization of the Resistome of Live Microbial Supplements
- Authors:
- Center:
-
Contributing OfficeCenter for Food Safety and Applied Nutrition
Abstract
Many dietary products available to the consumer, such as dietary supplements, contain live microorganisms purported to confer a human health benefit. The potential spread of antimicrobial resistance (AMR) from probiotic strains, present in commercial products, to commensal human gut microbiota is of serious concern to public health. The goal of this project is to determine the extent and content of AMR in commercial live microbial supplements using a culture-independent metagenomic approach. DNA extracted from the products was sequenced using the Illumina NextSeq platform. De novo assemblies were analyzed for the presence of AMR genes using the Resistance Gene Identifier, StarAMR, and AMRFinder tools available on GalaxyTrakr. The contigs shown to be positive for AMR genes were run on NCBI Blast to identify which bacterial species they were associated with. NCBI Blast can also determine the location of the AMR genes (chromosome or plasmid). For the AMR genes found to be located on the chromosome, the presence of mobile genetic elements, known to have a role in the dissemination of AMR within the microbial community, was determined. PHASTER (Phage Search Tool Enhanced Release) was used for identification and annotation of prophage sequences. MobileElementFinder was used to identify miniature inverted repeats (MITEs), insertion sequences, unit transposons, composite transposons, integrative mobilization elements (IMEs), integrative conjugative elements (ICEs), and plasmids. The position and distance of the mobile elements in relation to the AMR gene was determined using MobileElementFinder. This analytical pipeline has shown potential as a tool for the identification of chromosome and plasmid-associated AMR from commercially available live microbial supplements. Data generated from this analytical pipeline can provide insight on the safety of these microbial supplements widely available to the consumer. This research will increase FDA’s regulatory tools to determine the safety of live microbial supplements and will help protect consumer’s health.