2023 FDA Science Forum
Genomic Structure and Diversity of spv Virulence Plasmids and Hybrid MDR-spv Virulence Plasmids in Salmonella
- Authors:
- Center:
-
Contributing OfficeCenter for Veterinary Medicine
Abstract
Introduction
Here we evaluate the composition of 38 spv virulence operon encoding plasmids (spv-plasmids) recovered from Salmonella isolated from retail meats, food animal cecal samples and diseased animals in the US.
Purpose
The purpose of this study was to evaluate the genetic diversity and structure of spv-plasmids carried by multiple Salmonella serovars with diverse AMR profiles.
Methods
Closed sequences of 38 Salmonella spv-plasmids from 8 Salmonella serovars were evaluated for AMR genes, virulence genes and plasmid Inc type using AMRFinder-plus and PlasmidFinder. Plasmids were grouped by genomic similarity through hierarchical clustering of plasmid pangenomic loci identified by Roary and piggy.
Results
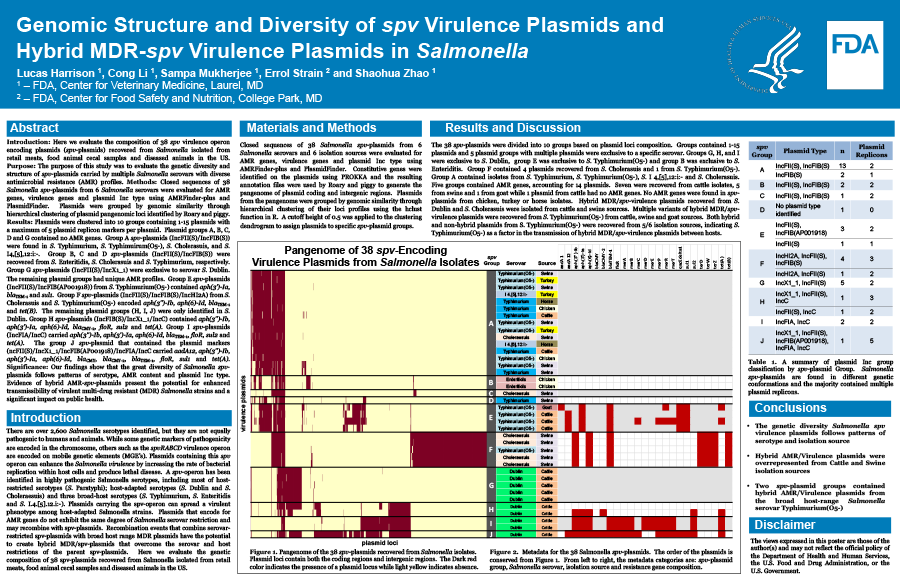
Plasmids were clustered into 10 groups containing 1-15 plasmids with a maximum of 5 plasmid replicon markers per plasmid. Plasmid groups A, B, C, D and G contained no AMR genes. Group A spv-plasmids (IncFII(S)/IncFIB(S)) were found in Typhimurium, Typhimuirum(O5-), Cholerasuis, and I4,[5],12:i:-. Group B, C and D spv-plasmids (IncFII(S)/IncFIB(S)) were recovered from Enteritidis, Cholerasuis and Typhimurium, respectively. Group G spv-plasmids (IncFII(S)/IncX1_1) were exclusive to serovar Dublin. The remaining plasmid groups had unique AMR profiles. Group E spv-plasmids (IncFII(S)/IncFIB(AP001918)) from Typhimurium(O5-) contained aph(3')-Ia, blaTEM-1 and sul1. Group F spv-plasmids (IncFII(S)/IncFIB(S)/IncHI2A) from Cholerasuis and Typhimurium(O5-) encoded aph(3'')-Ib, aph(6)-Id, blaTEM-1 and tet(B). The remaining plasmid groups (H, I, J) were only identified in S. Dublin. Group H spv-plasmids (IncFIB(S)/IncX1_1/IncC) contained aph(3'')-Ib, aph(3')-Ia, aph(6)-Id, blaCMY-2, floR, sul2 and tet(A). Group I spv-plasmids (IncFIA/IncC) carried aph(3'')-Ib, aph(3')-Ia, aph(6)-Id, blaTEM-1, floR, sul2 and tet(A). The group J spv-plasmid (IncFII(S)/IncX1_1/IncFIB(AP001918)/IncFIA/IncC) carried aadA12, aph(3'')-Ib, aph(3')-Ia, aph(6)-Id, blaCMY, blaCMY-2, blaTEM-1, floR, sul1 and tet(A).
Significance
Our findings show that the great diversity of Salmonella spv-plasmids follows patterns of serotype, AMR content and plasmid Inc type. Evidence of hybrid AMR-spv-plasmids present the potential for enhanced transmissibility of virulent MDR Salmonella strains and a significant impact on public health.