2023 FDA Science Forum
Genomic Structure and Diversity of Plasmids in Campylobacter coli and C. jejuni
- Authors:
- Center:
-
Contributing OfficeCenter for Veterinary Medicine
Abstract
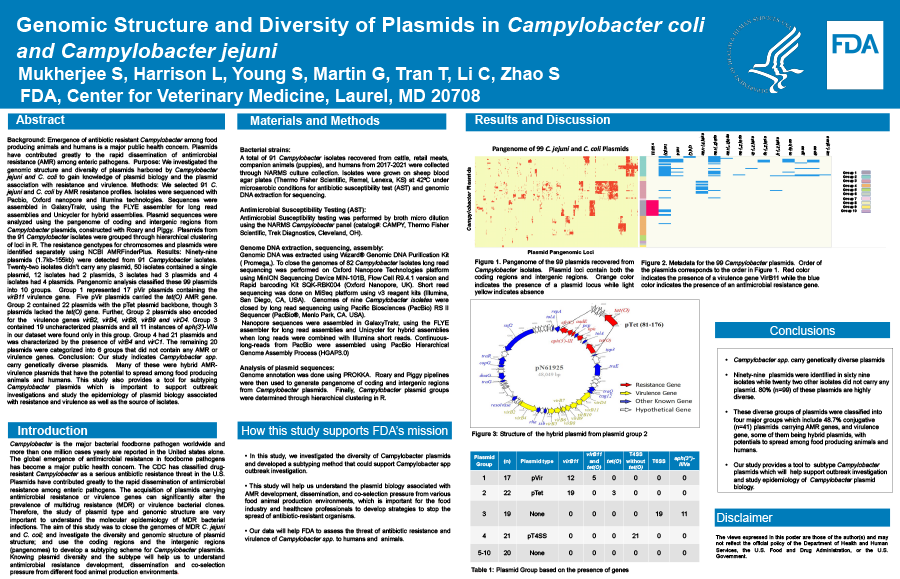
Background: Emergence of antibiotic resistant Campylobacter among food producing animals and humans is a major public health concern. Plasmids have contributed greatly to the rapid dissemination of antimicrobial resistance among enteric pathogens. Purpose: We investigated the genomic structure and diversity of plasmids harbored by Campylobacter jejuni and C. coli to gain knowledge of plasmid biology and the plasmid association with resistance and virulence. Methods: We selected 91 C. jejuni and C. coli by AMR resistance profiles. Isolates were sequenced with Pacbio, Oxford Nanopore and Illumina technologies. Sequences were assembled in GalaxyTrakr, using the FLYE assembler for long read assemblies and Unicycler for hybrid assemblies. Plasmid sequences were analyzed using the pangenome of coding and intergenic regions from Campylobacter plasmids, constructed with Roary and Piggy. Plasmids from the 91 Campylobacter isolates were grouped through hierarchical clustering of loci in R. Resistance genotypes for chromosomes and plasmids were identified separately using NCBI AMRFinderPlus. Results: Ninety-nine plasmids (1.7kb-155kb) were detected from 91 Campylobacter isolates. Twenty-two isolates harbored no plasmids, 50 isolates contained a single plasmid, 12 isolates had 2 plasmids, 3 isolates had 3 plasmids and 4 isolates had 4 plasmids. Pangenomic analysis classified these 99 plasmids into 10 groups. Group 1 represented 17 pVir plasmids containing the virB11 virulence gene. Five pVir plasmids carried the tet(O) AMR gene. Group 2 contained 22 plasmids with the pTet plasmid backbone, though 3 plasmids lacked the tet(O) gene. Group 2 plasmids also encoded for the virulence genes virB2, virB4, virB8, virB9 and virD4. Group 3 contained 19 uncharacterized plasmids and was the only group to encode for aph(3')-VIIa. Group 4 had 21 plasmids and was characterized by virB4 and virC1. The remaining 21 plasmids were categorized into 6 groups containing neither AMR nor virulence genes. Conclusion: Our study indicates Campylobacter spp. carry genetically diverse plasmids. Many plasmids were hybrid AMR-virulence plasmids that have the potential to spread among food producing animals and humans. This study also provides a tool for subtyping Campylobacter plasmids to support outbreak investigations and study the epidemiology of plasmid biology associated with resistance, virulence and the source of isolates.