2021 FDA Science Forum
Forced Degradation Study of Rituximab Using the Multi-Attribute Method (MAM)
- Authors:
- Center:
-
Contributing OfficeCenter for Drug Evaluation and Research
Abstract
Background
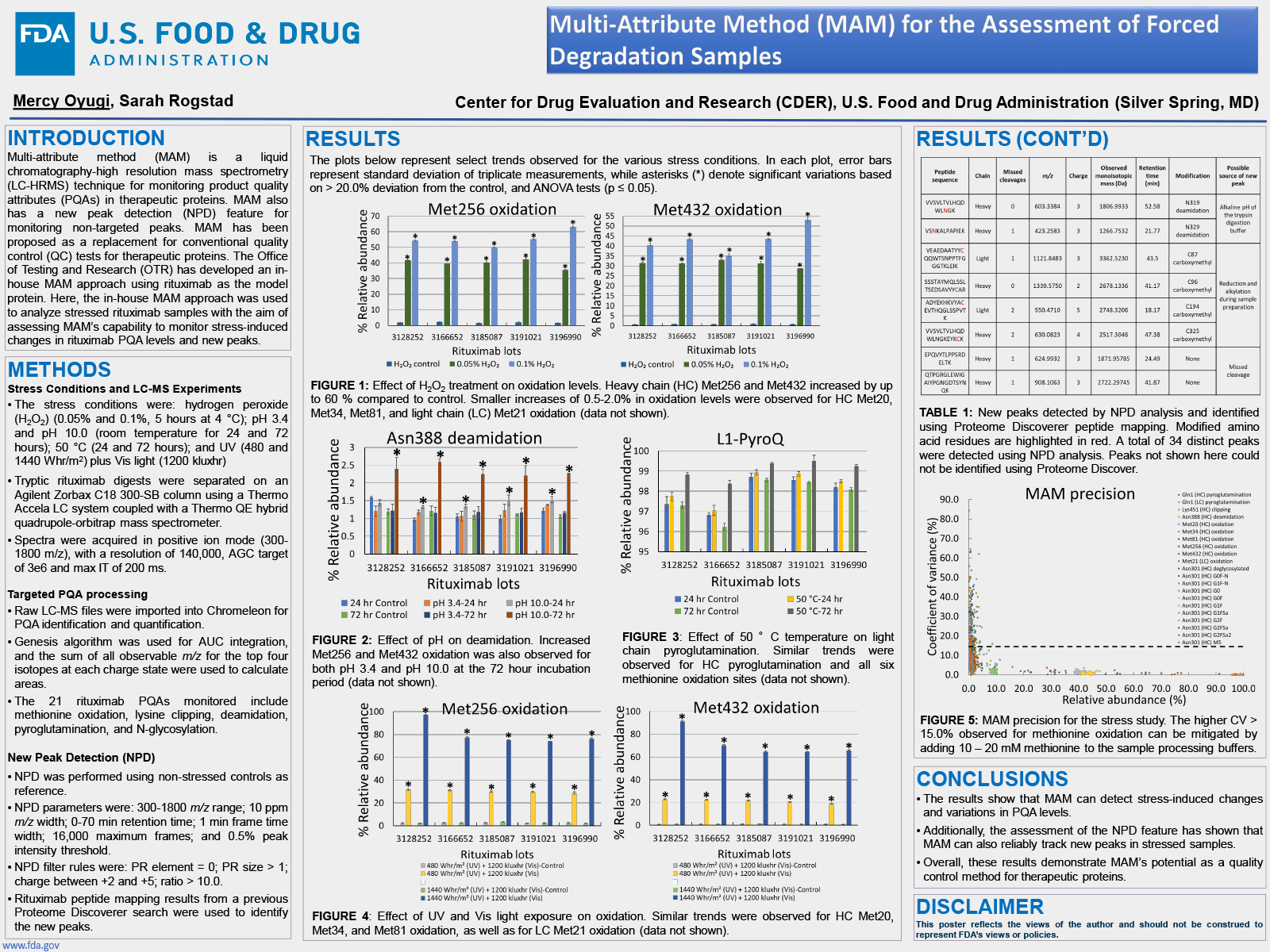
Multi-attribute method (MAM) is a liquid chromatography-high resolution mass spectrometry (LC-HRMS) technique for monitoring product quality attributes (PQAs) in therapeutic proteins. MAM also has a new peak detection (NPD) feature for monitoring impurities. MAM has been proposed as a replacement for conventional quality control (QC) and release methods for therapeutic proteins. The Office of Testing and Research (OTR) has developed an in-house MAM approach using rituximab as a model protein, which was used in this study to analyze rituximab samples after exposure to various stress conditions.
Purpose
To assess MAM’s capability to monitor stress-induced changes in rituximab PQA and impurity levels.
Methods
Rituximab samples were exposed to the following stress conditions: hydrogen peroxide (H2O2) (0.05% and 0.1% incubated for 5 hours at 4 °C); pH 3.4 and pH 10.0 (incubated at room temperature for 24 and 72 hours); 50 °C (incubated for 24 and 72 hours); and UV light (480 Whr/m2 and 1440 Whr/m2) applied simultaneously with Vis light (1200 kluxhr). Non-treated controls were incubated along with the stressed samples. The samples were then digested using trypsin, followed by LC-HRMS runs of the digests. Data analysis was then performed using Chromeleon software.
Results
The 21 rituximab PQAs quantified included two pyroglutamination sites, Asn388 deamidation, Lys451 clipping, six methionine oxidation sites, and 11 N-glycans. Variations in PQA levels were considered significant if the ANOVA p-value was ? 0.05 and the relative abundance deviated by > 20.0% from the control. Except for pH 3.4, all stress conditions caused significant increases in oxidation levels, with Met256 and Met432 being the most affected. A significant increase in Asn388 deamidation was observed at pH 10.0, while increased pyroglutamination was observed at 50 °C. None of the stress conditions significantly affected Lys451 clipping or N-glycan levels. For NPD, a total of 100 new peaks were detected across all samples. 46/100 peaks were identified using Proteome Discoverer, with 25 peaks being artifacts of sample preparation steps.
Conclusion
Overall, the results show that MAM can detect stress-induced variations in PQA levels while also monitoring impurities, thus demonstrating potential for its application in QC of therapeutic proteins.