2021 FDA Science Forum
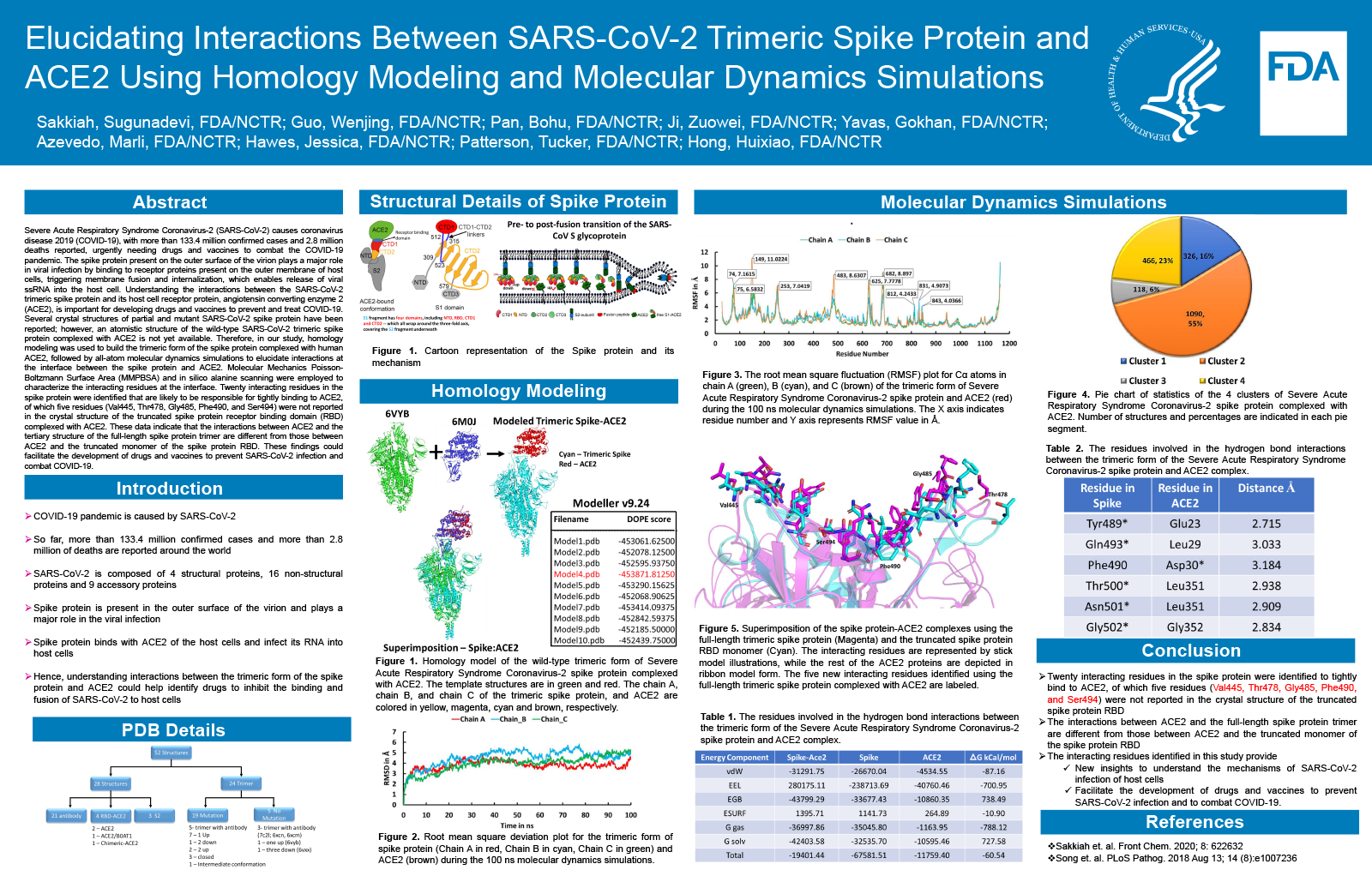
Elucidating Interactions Between SARS-CoV-2 Trimeric Spike Protein and ACE2 Using Homology Modeling and Molecular Dynamics Simulations
- Authors:
- Center:
-
Contributing OfficeNational Center for Toxicological Research
Abstract
Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) causes coronavirus disease 2019 (COVID-19), with more than 100.4 million confirmed cases and 2.1 million deaths reported, making drug and vaccine development to combat COVID-19 immensely important. The spike protein present on the outer surface of the virion plays a major role in viral infection by binding to receptor proteins present on the outer membrane of host cells, triggering membrane fusion and internalization, which enables release of viral ssRNA into the host cell. Understanding the interactions between the SARS-CoV-2 trimeric spike protein and its host cell receptor protein, angiotensin converting enzyme 2 (ACE2), is important for developing drugs and vaccines to prevent and treat COVID-19. Several crystal structures of partial and mutant SARS-CoV-2 spike proteins have been reported; however, an atomistic structure of the wild-type SARS-CoV-2 trimeric spike protein complexed with ACE2 is not yet available. Therefore, in our study, homology modeling was used to build the trimeric form of the spike protein complexed with human ACE2, followed by all-atom molecular dynamics simulations to elucidate interactions at the interface between the spike protein and ACE2. Molecular Mechanics Poisson-Boltzmann Surface Area (MMPBSA) and in silico alanine scanning were employed to characterize the interacting residues at the interface. Twenty interacting residues in the spike protein were identified that are likely to be responsible for tightly binding to ACE2, of which five residues (Val445, Thr478, Gly485, Phe490, and Ser494) were not reported in the crystal structure of the truncated spike protein receptor binding domain (RBD) complexed with ACE2. These data indicate that the interactions between ACE2 and the tertiary structure of the full-length spike protein trimer are different from those between ACE2 and the truncated monomer of the spike protein RBD. These findings could facilitate the development of drugs and vaccines to prevent SARS-CoV-2 infection and combat COVID-19.