A New Approach to Pharmacometrics: Recurrent Neural Networks for Modeling Drug Exposure and Drug Response
Pharmacometric Models as Tools in Drug Development and Review
Pharmacokinetic (PK) models are increasingly relied upon in drug development and review. Typically consisting of a set of linked differential equations, these models allow one to relate the dose given, patient characteristics, and the time(s) of administration with temporal changes in drug concentrations in the body. The models’ equations incorporate available biologic information (e.g., anatomy, physiology, and biochemistry) as well as known values pertaining to specified drug substance (e.g., rates of dissolution, metabolism, and transport). In model development, these parameters and the equations themselves may be adjusted so that the model predictions fit the available empirical data as closely as possible. Using similar mathematical approaches, pharmacodynamic (PD) models try to capture the temporal effects of a drug (for example, in inhibiting an enzyme or changing expression of a given protein in the body). A combined PK/PD model provides both the predicted changes in the drug concentrations in specific bodily compartments over time and the time course of one or more biologic effects that result from these concentration changes.
Pharmacometric models are critically important because when validated against independent empirical data they allow us to better understand drug effects in individuals and populations who are not included in clinical trials and to explore additional real-world scenarios. For example, a model that incorporated information about the impact of the kidney or liver function on the clearance of a certain drug could be used to reasonably predict drug exposure in patients experiencing kidney impairment or prone to drug-drug interactions. By pursuing research into pharmacometric models and by advancing a general framework for assessing their credibility, CDER has been able to offer guidance to sponsors on the use of modeling approaches, thereby fostering drug development broadly and promoting an increase in the number of regulatory submissions that use pharmacometric approaches.
Constructing Pharmacometric Models Using Artificial Neural Networks
Model development can be challenging, owing to the highly complex nature of the physiological mechanisms that govern time-dependent changes of drug concentrations in an individual patient and their biological effects. CDER researchers have recently investigated how methods that use artificial neural networks (some basic elements of these are shown in Figure 1) can be applied to modeling problems.1 Specifically, CDER scientists have developed a model based on a kind of recurrent neural network to simulate the time course of a PD response that is not directly related to the drug concentration, but rather that develops latently, according to complex biological intermediate steps. To test the capability of machine learning to model PD in this scenario, the researchers first constructed a mechanistic PK/PD model for a hypothetical drug for which there was a delayed biological response (a change in the concentration of a biomarker) to changes in drug concentration. Using this model, they generated simulated data (both PK and PD profiles) for patients differing by demographics and weight to whom the drug was given daily over a period of seven days.
Figure 1. Artificial Neural Networks. The building block of an artificial neural network is the single neuron (or perceptron) (left panel). Numerical inputs (x) from a data set are multiplied by individual weights (w) and then these products are added together along with a numerical bias term. A mathematical “activation” function converts this sum into a number between 0 and 1 (the neuron’s output). Individual neurons are combined into neural networks (right panel). In the simple “feed forward” network shown, information is propagated to the output layer according to weights and bias factors as in the left panel. The numerical values arriving in this output layer can be compared against a correct solution to compute the prediction error. The individual weights of the network are then adjusted such that the network generates a new output that is closer to the desired solution. In training the network to complete a task, the process of forward and backward propagation of information stops when the error appears to have been minimized.
There are many variations in the architecture of neural networks. CDER researchers constructed a long short-term memory recurrent neural network and described its architecture. This type of network has the property that it can retain and use information from much earlier states of the network and is especially suited to tasks in which the data are sequential (for example language or a patient’s pharmacologic response to a drug).
The researchers chose to develop a type of recurrent neural network (RNN) called a long short-term memory recurrent neural network (LSTM RNN), because it had shown great promise in predicting sentences and other kinds of sequential data. They constructed several LSTM RNN models that varied in their number of hidden layers and perceptrons and trained them to use the time sequence of plasma concentrations simulated by their mechanistic model along with patient baseline PD values and patient demographics to predict the PD response under one specific dosing schedule. The simplest model that predicted this PD response with acceptable error was then used to predict PD response in patients given the drug according to different schedules.
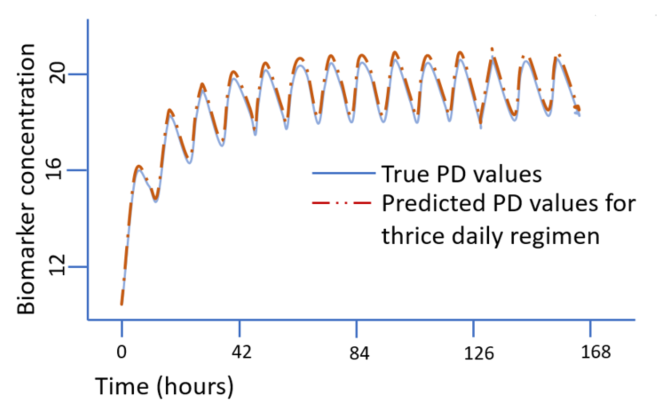
Figure 2. CDER researchers constructed a kind of recurrent neural network that has proved valuable for language prediction and other tasks where sequence is important, i.e., a long short-term memory recurrent neural network (LSTM RNN). When trained on simulated data from a traditional PK/PD model for patients given a hypothetical drug once a day, this neural network learned to predict PD response in these patients. Notably, the same trained model could predict response when hypothetical patients were placed on a different dosing regimen. The traces show the “true” PD response based on the researchers’ PK/PD model and the neural network’s prediction of the appearance of the biomarker (PD response) for a thrice-daily dosing regimen.
The machine learning model was able to accurately predict the PD response of individuals who were treated with dosing regiments that were different from the once-daily regimen used in model construction (Figure 2). This finding is significant, because dosing regimens are often changed during a drug’s development process. The authors also tested the LSTM RNN model when it was trained with sparse data. It could still predict PD response when the dosing regimen was the same as that used for training (once daily) but could not do so accurately for patients on twice- or thrice-daily regimens.
In their report, the researchers indicated that model prediction with unevenly distributed time points is an active area of research in machine learning and that another challenge would be to predict more complex PD responses, such as responses in which there were more layers of delay between PK and PD response, and highly complex interactions among biomarkers.
The CDER investigators suggest that when scientists from the machine learning and mechanistic modeling disciplines interact and learn from each other, they will find many overlapping concepts, equations, and ways to visualize model structure architectures, and that machine learning models and more traditional mechanistic ones may complement each other to improve the efficiency of drug development and optimize treatments for individual patients.
How does this work advance drug development and evaluation?
This study demonstrated that simulated pharmacokinetic/pharmacodynamic data can be analyzed with a machine learning algorithm. This finding suggests that with further research, neural network-based models such as LSTM RNNs may complement traditional PK/PD models in the area of highly complex PK/PD data analysis and possibly facilitate development of predictive models with improved accuracy.
- 1Liu, Xiangyu, et al. "Long short-term memory recurrent neural network for pharmacokinetic-pharmacodynamic modeling." International Journal of Clinical Pharmacology and Therapeutics (2020).